PNAS | University of Chinese Academy of Sciences (Hangzhou) Chen Luonan Group and MOMED BIOTECH Mine Lysine Post-Translational Modification Sites by Integrating Protein Language Models with Structural Features
Good Fortune in the Year of the Horse · Warm Congratulations
The team of Chen Luonan from the Hangzhou Institute for Advanced Study, University of Chinese Academy of Sciences (UCAS) / Shanghai Jiao Tong University, in collaboration with MOMED BIOTECH, recently published their latest research findings in the journal PNAS.
Recently, the research team led by Chen Luonan from the Hangzhou Institute for Advanced Study, University of Chinese Academy of Sciences (UCAS) / Shanghai Jiao Tong University, in collaboration with MOMED BIOTECH, published an online research paper in the journal PNAS titled "Mining Lysine Post-Translational Modification Site by Integrating Protein Language Model Representations with Structural Context." This study proposes a deep learning framework that integrates protein language models with atomic-level structural features, achieving accurate prediction of various lysine post-translational modification (PTM) sites. The functional impact of these sites was further validated through molecular dynamics simulations, offering new insights for PTM-related mechanistic research and drug target development. The co-first authors are Dr. Luo Mengqi from the Hangzhou Institute for Advanced Study, UCAS, and Dr. Zhu Xiaohong from MOMED BIOTECH. The co-corresponding authors are Dr. Chen Luonan from the Hangzhou Institute for Advanced Study, UCAS / Shanghai Jiao Tong University, Dr. Warshel from the University of Southern California / MOMED BIOTECH, and Dr. Bai Chen.
Lysine PTMs are critical molecular events that regulate protein function, signal transduction, and disease progression. However, their experimental identification still faces challenges such as high costs and low throughput. Most existing computational methods rely on manually designed sequence features, struggle to effectively integrate three-dimensional structural information, and are often limited to a single PTM type, resulting in poor generalization. Constructing an intelligent prediction model capable of integrating multi-source information and uniformly mining multiple PTM types has become a bottleneck that needs to be broken in this field.
This study proposes a dual-module deep learning framework. On one hand, it utilizes the protein language model ESM-2 to extract semantic features from sequences. On the other hand, based on structural data provided by AlphaFold, it constructs atomic-level contact maps and captures spatial structural information through a Graph Convolutional Network (GCN). After fusing these two types of features, PTM site classification is achieved via a Multilayer Perceptron. This framework demonstrates stable and excellent predictive performance across six common types of lysine PTMs (including acetylation, succinylation, crotonylation, etc.), with F1-scores reaching up to 80.9% and AUC up to 88.3%. It maintains strong generalization ability under different data partitioning strategies, with a small fluctuation range in results upon repeated evaluation (±1.0–1.5%).
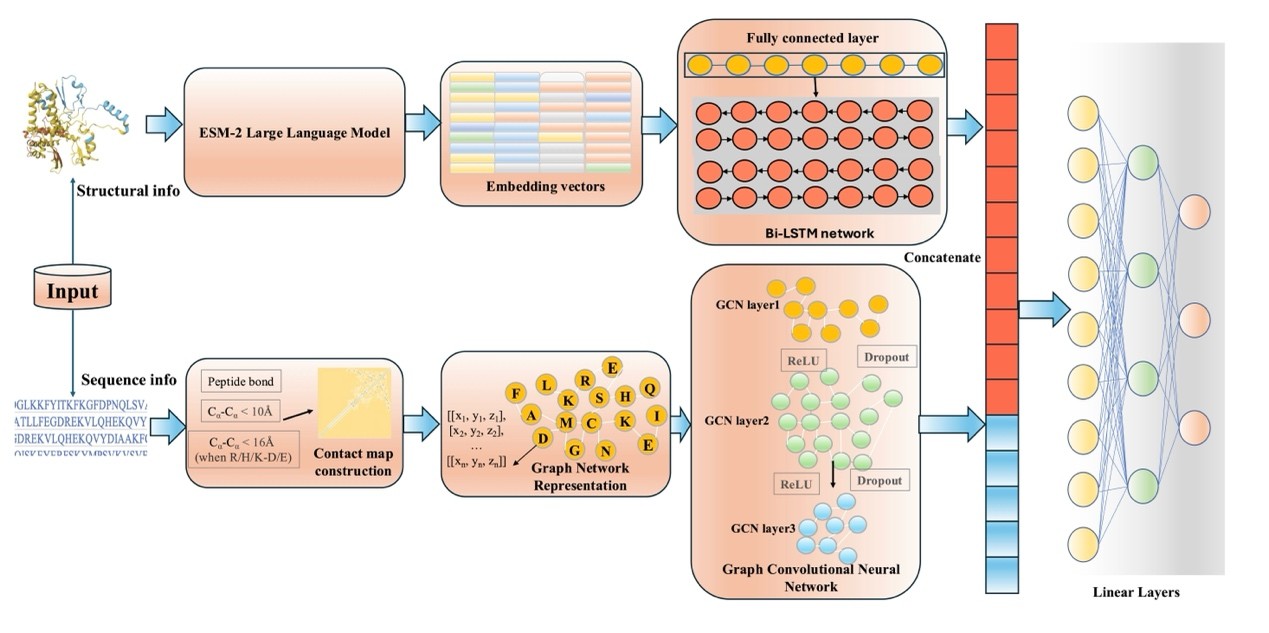
Figure 1. Overall architecture of the prediction model. The model primarily consists of a structural information processing module and a sequence information processing module. The feature vectors output by these two modules are concatenated and then fed into a fully connected network for dimensionality reduction to generate the final output. Specifically, the structural information processing module constructs a contact map based on the atomic-level three-dimensional coordinates of amino acids and analyzes it through a graph neural network. The sequence information processing module utilizes a large language model to obtain representations, which are then further processed through a linear layer and a bidirectional long short-term memory network.
To further validate the practical application value of the model, the research team applied it to predict PTM sites on human C-type lectin domain family 12 member A (hCLEC12A). The analysis identified K181 and K174 as potential acetylation/crotonylation modification sites. Subsequently, using all-atom molecular dynamics simulations, the team systematically compared the binding modes and changes in binding free energy between unmodified and modified hCLEC12A when interacting with the antibody 50C1. The results indicated that the modified systems (particularly K181 acetylation and the dual modification system) significantly weakened the protein-antibody interaction. The energy contributions of key interfacial residues were markedly altered, consequently affecting the stability of the complex.
Figure 2. Impact of hCLEC12A post-translational modifications on 50C1 antibody recognition. (A) Binding interface of 50C1 with hCLEC12A under (A) K181 acetylation or (B) K174 crotonylation combined with K181 acetylation. Hydrogen bonds, salt bridges, and π-π stacking interactions are indicated by yellow, cyan, and purple dashed lines, respectively. Changes in per-residue free energy decomposition of key hCLEC12A residues under (C) K181 acetylation and (D) K174 crotonylation combined with K181 acetylation conditions, relative to the unmodified system. ∆∆G_decomp = ∆G_decomp (modified system) – ∆G_decomp (unmodified system). Data are presented as mean ± standard error. A more negative value indicates a greater contribution of that amino acid to the binding between hCLEC12A and 50C1.
This study not only proposes a unified prediction framework scalable to multiple PTM types but also integrates AI predictions with biological function validation through dynamic simulations, overcoming the limitations of traditional static prediction methods. This "prediction-validation" closed-loop process provides important tools and a theoretical basis for future PTM functional exploration, disease mechanism analysis, and the design of targeted drugs based on key modification sites.
Interested readers can access the original research paper at:https://www.pnas.org/doi/10.1073/pnas.2529141123
The code and data associated with this article have been made publicly available on the GitHub platform(https://github.com/qi29/lysine-PTM-site-Mining).
END
NEW YEAR -
Hot News
-
2022-11-05
The Bai Chen Research Group Publishes Paper in the International Journal of Molecular Sciences
- 2022-12-20
- 2022-12-26
- 2026-02-25




